Molecular Genetic Identification of Some Sweet Sorghum - Sorghum bicolor L. (Ankolib) Accessions - Sudan
Received: 16-Jan-2023, Manuscript No. PULJMBR-23-6057; Editor assigned: 18-Jan-2023, Pre QC No. PULJMBR-23-6057 (PQ); Accepted Date: Jan 25, 2023; Reviewed: 20-Jan-2023 QC No. PULJMBR-23-6057 (Q); Revised: 22-Jan-2023, Manuscript No. PULJMBR-23-6057 (R); Published: 27-Jan-2023, DOI: 10.37532/puljmbr.2023.6(1).5-13
Citation: Ahmed AA, Hassaballa LA, Molecular genetic identification of some sweet sorghum-Sorghum Bicolor L. (Ankolib) accessions-Sudan. J Mic Bio Rep. 2023;6(1):5-13
This open-access article is distributed under the terms of the Creative Commons Attribution Non-Commercial License (CC BY-NC) (http://creativecommons.org/licenses/by-nc/4.0/), which permits reuse, distribution and reproduction of the article, provided that the original work is properly cited and the reuse is restricted to noncommercial purposes. For commercial reuse, contact reprints@pulsus.com
Abstract
This study is an attempt to identify some Sudanese Sweet Sorghum (Ankolib) [Sorghum bicolor (L.) Moench.] accessions using two types of DNAbased markers: RAPD and SS. Seven (Ankolib) accessions were assayed, namely: Black Ankolib, Black White Ankolib, Dark Red Ankolib, Red Yellow Ankolib, White Ankolib, White Black Ankolib and Bengaga. All of the accessions were uniquely identified and fingerprinted. The levels of polymorphism among the accessions as revealed by 22 RAPD primers and 16 SSR primer pairs were (58%) and (76%) respectively, indicating that SSRs markers were highly polymorphic. The scored data of the two markers were analyzed using the Dice Coefficient to assess genetic relationships among the 7 (Ankolib) accessions. The results of the statistical analysis revealed that the accession Bengaga was distantly related to the other 6 accessions which all showed a close genetic similarity among them. UPGMA cluster analysis generated a dendrogram for each marker alone and for the combined data of the two markers. It was observed that in all of the dendrogram the accession Bengaga was found in a unique cluster thus indicating its uniqueness, since it is the only accession that has seeds that produce flour besides its juicy sweet stem.
Keywords
Ankolib; Sweet sorghum; Molecular genetics
Introduction
The classification and identification of all known plants have gradually shifted from purely artificial systems to more accurate systems that are based on genetic experimentations. This development of Plant Taxonomy is a reflection of the development and application of new techniques. Taxonomists have now made new techniques applied in the field of Ecology, Histology, Physiology, Biochemistry, Cytogenetic and Molecular Biology. The accurate, fast, reliable, and cost-effective identification of plant populations and varieties is essential in agriculture as well as in pure and applied plant research. Methods of varietals identification are most frequently based on an assessment of a range of morphological characteristics, while these methods are very effective for many purposes, the morphological comparison may have limitations including subjectivity in the analysis of the character, the influence of environmental or management practices on the character limited diversity among cultivars with highly similar pedigrees and the confining of expression of some diagnostic characters to a particular stage of development, such as flowering and fruit ripening. These considerations have led to the exploration or adoption of other techniques for varietals identification [1]. New techniques based on DNA profiling provide novel approaches to varietals identification which offer advantages over all other techniques and methods for demonstrating distinctness because the DNA sequence of an organism is unique and independent of environmental conditions or management practices. The presence of the same DNA in every living cell of the plant allows tests of any tissue at any stage of growth (provided that the DNA is sufficient purity). The recent advent of the polymerase chain reaction PCR has enabled the development of new DNA profiling techniques that are simply and quickly performed. Therefore, attention has increasingly focused on the DNA molecule as a source of informative polymorphisms. DNA profiling techniques have two applications:
• DNA Fingerprinting identifies individuals by the unique profile produced when their DNA is separated into a series of fragments and resolved into sized classes. This technique has potential in the specific identification of plant cultivars and their subsequent protection.
• Gene Mapping allows researchers to mark genes or chromosome regions that are related to genetic traits such as pests or disease resistance, fruit size, fruit color and stress- tolerance…etc
A particular advantage of such a technique is that complex polygenic trait can be analyzed. Mapping techniques can also be used to isolate genes based entirely on their behavior. Once these have been identified, sequenced, and cloned. Gene transfer techniques can be used to transfer them to other species. The sustainable development of large areas of the world is today one of the greatest challenges. The living conditions of large parts of the world population are greatly affected by the availability of energy, together with food and water. As most developing countries have limited fossil fuel indigenous resources, the import of energy fills the increasing difference between demand and production. Consequently, nowadays a large number of motivations and a growing interest exist for the development of renewable sources of energy, like energy systems based on biomass, in particular energy crops, such as sweet sorghum.
Sweet sorghum has been selected as the main biomass resource for the following reasons:
• It has a very high yield (up to 80 tons/ha. of fresh matter) in terms of starch, sugar, and lignocellulosic component and today can be considered the most versatile and promising crop (food-feedenergy).
• It has two times the photosynthetic efficiency in comparison with Sugar Beet, Soybean, Wheat…etc., (Its photorespiration can almost not be measured).
• It can be grown in a wide range of latitudes (tropical, sub-tropical, and temperate zones) as well as on poor-quality soil.
• It requires very low fertilizer and water input. The water requirement for growing sweet sorghum is one of the lowest: about 1/2 of the requirement for corn and about 1/3 of the requirement for sugar cane.
• Its growth period is short (4 months -5 months) in comparison with Sugar cane (8 months-24 months). The versatility of sweet sorghum is reflected in the range of research going on around the world into sweet sorghum production.
• It is a low water-use, stress-tolerant crop with considerable potential for future small and large-scale exploitation. All these benefits and the importance of the sweet sorghum as the most promising (foodfeed-energy) crop for the twenty-first century and its potential to aid development to encourage the author to carry out this study In this study, we are going to identify some sweet Sorghum bicolor (L.) Moench. accessions in Sudan, using DNA-based markers like random amplified polymorphic DNA (RAPD) and Simple Sequence Repeats (SSRs).
Scientific Classification
Sweet Sorghum bicolor (L.) Moench. which represents the scope of this study, belongs to the family Gramineae (Poaceae). It can be classified as follows: (Table 1).
TABLE 1 Scientific Classification
| Classification | |
|---|---|
| Kingdom | Plantae |
| Subkingdom | Tracheobionta |
| Superdivision | Spermatophyta |
| Division | Magnoliophyta |
| Class | Liliopsida |
| Subclass | Commelinidae |
| Order | Cyperales (Poales) |
| Family | Gramineae (Poaceae) |
| Genus | Sorghum Moench. |
| Species | Sorghum bicolor (L.) Moench |
Botanical description
Sorghum is a vigorous grass that varies between (0.5 m and-5.0 m) in height. It is an annual plant. It produces one or many tillers, which emerge initially from the base and later from stem nodes. The root system consists of fibrous adventitious roots that emerge from the lowest nodes of the stem, below and immediately above ground level. Roots are normally concentrated in the top (0.9 m) of soil but may extend to twice that depth and can extend to (1.5 m) in the lateral spread. The stem is solid, usually erect. Its center can be dry or juicy, insipid or sweet to taste. The center of the stem can become pithy with spaces. Leaves vary in number from 6–24, depending on the cultivar. They are borne alternately in two ranks. Leaf sheaths vary in length between (15 cm-35 cm) and encircle the stem with their margins overlapping. The leaf sheath often has a waxy bloom. Leaves are from (30 cm-135 cm) long and (1.5 cm-13 cm) wide, with flat or wavy margins. Midribs are white or yellow in dry pithy cultivars or green in juicy cultivars. The inflorescence is a panicle, usually erect, but sometimes recurved to form a gooseneck. The panicle has a central rachis, with short or long primary, secondary, and sometimes tertiary branches, which bear groups of spikelets. The length and closeness of the panicle branches determine panicle shape, which varies from densely packed conical or oval to spreading and lax. The grain is usually partially covered by glumes. The seed is rounded and bluntly pointed, from (4 cm-8 mm) in diameter, and varies in size, shape, and color with cultivar.
Geographic distribution
Sorghum originated in the northeastern quadrant of Africa, where the greatest variability in wild and cultivated species is found to this day. It was probably domesticated in Ethiopia by selection from wild sorghum between 5.000 years and 7.000 years ago. From this center of origin, it was distributed along trade and shipping routes throughout Africa, and through the Middle East to India at least 3.000 years ago. It reached China along the silk route. Sorghum was first taken to America through the slave trade from West Africa. It was reintroduced in the late 19th century for commercial cultivation and has subsequently been introduced into South America and Australia. Although wild varieties of sorghum are attested as early as 8.000 years ago in the Nilotic regions of southern Egypt and Sudan, the location of its true domestication within East Africa is still speculative. It is widely held that the genetic separation of domesticated Sorghum bicolor L. from its progenitor did not occur much before 2.000 years ago somewhere in East Africa, possibly the Ethiopian highlands, but more likely further west. The presence of true domesticated S. bicolor L. is claimed much earlier than this (3.700 years–4.900 years ago in India, Oman, and Yemen, although the identity of the remains as full domesticates is still disputed. Sorghum is now widely found in the drier areas of Africa, Asia, America, and Australia.
Uses of Sorghum bicolor (L.)
Sweet Sorghum bicolor (L.) Moench. is the only crop that provides grain and stem that can be used for sugar, alcohol, syrup, jaggery, fodder, fuel, bedding, roofing, fencing, paper, and chewing [2]. Sweet sorghum has been widely used for the production of forage and silage for animal feed. It has also been used for nearly 150 years to produce concentrated syrup with the distinctive flavor mentioned, the primary products from sweet sorghum are based on exploiting the high levels of sugars found in the stem [3,4]. These products include ethanol, Syrup, crystalline sugar, electricity produced from burning the residual fiber and Particle board or paper (from the fiber). Sucrose–a type of sweet sorghum, which mainly contains sucrose, can be used for refining crystal sugar. Syrup-type sweet sorghum, which mainly contains glucose, can be used for producing syrup; it is also a material of quality for making drinking wine and alcohol also noted that sweet sorghum has a medicinal use. The inflorescence has astringent, hemostatic, and antidiarrhea properties and can be administered as an infusion, tincture and medicinal wine [4]. The stem is used in Nigerian traditional medicine for the treatment of anemia, especially during pregnancy. The oil crisis of 1973 and 1976 renewed interest in the commercial production of sweet sorghum for biological transformation into ethyl alcohol for use as fuel or fuel additive [5]. In Sudan, grain sorghum contributes about (39%) of the calories in the human diet, while sweet sorghum is mainly used as fodder. In Sinnar locality, the grains are ground and used as special flour.
Molecular studies
Recent advances in Molecular Biology, principally in the development of the Polymerase Chain Reaction (PCR) for amplifying DNA, DNA sequencing, and data analysis have resulted in powerful techniques which can be used for the screening, characterization, and evaluation of genetic diversity. The extensive number of research articles currently appearing in the literature, describing the use of these techniques in a wide range of plant species and diversity problems, is testimony to their increasing impact in this field. The application of DNA technology in seed research has progressed rapidly during the last ten years, especially in the area of cultivar identification. DNA fingerprinting technique has a promising application in this respect. A cultivar identification test based on DNA genetic fingerprints would be much faster than the official methods of variety identification, which require the growing of suspect plants to the flowering stage, in parallel with control plants reported that the use of DNA-based markers for the genetic analysis and manipulation of important agronomic traits has become an increasingly useful tool in plant breeding [6]. DNA markers have the potential to enhance the operation of plant breeding programs through several ways ranging from genetic diversity, increasing the efficiency of selection for difficult traits, and making environmentally neutral selection possible. The problem of DNA extraction is still an important issue in the field of plant molecular biology. A good extraction procedure for the isolation of DNA should yield adequate and intact DNA of reasonable purity. The procedure should also be quick, simple, and cheap. DNA extraction is difficult in a variety of plants because of the presence of metabolites that interfere with DNA isolation procedures and downstream applications such as DNA restriction, amplification, and cloning [7]. Various plants contain high levels of polysaccharides and many types of secondary metabolites affecting DNA purification. A good protocol for the isolation of DNA from leaf material containing large quantities of polyphenols, tannins, and polysaccharides was reported by using a modified CTAB extraction method by applying high salt concentrations to remove polysaccharides and PVP to remove polyphenols [8]. This modified CTAB protocol was used for DNA extraction from dry and fresh leaves of pearl millet (Pennisetum glaucum), they mentioned that this method solved the problems of DNA degradation, contamination, and low yield due to binding and/or co-precipitation with starches and polysaccharides reported that, the knowledge of genetic relationships and distance among individuals provide useful information for breeders, and this can then be exploited to obtain the benefit from the potential of each population [9]. They also stated that molecular-assisted genetic analysis provides means to locate and select genes controlling important agronomic, pest-resistance, stress-tolerance, and food quality traits. Genetic relationships facilitate the identification of diverse parents to cross in hybrid combinations to maximize the expression of heterosis [10-12]. India reported that two sweet sorghum varieties (S21-3-1 and S23-1-1) were developed by crossing American lines with a local variety [13]. These two varieties were found to be useful for their sugar content to produce ethanol and their grains to produce flour. Analysis of the extent and distribution of genetic variation in a crop is essential in understanding the evolutionary relationship between accession and sampling genetic resources more systematically for breeding and conservation mentioned that accurate identification of new varieties is essential to plant variety protection [14,15]. The Plant Variety Protection Act (PVPA) of (1970) was established to help plant breeders protect their crop cultivars from commercial exploitation by others. DNA-based markers played a great role in this respect since they provide unique DNA fingerprints for plant varieties and cultivars ensuring their protection under the (PVPA). Various techniques have been used to mark individual characters in segregating populations and constructing genetic maps. Initially, isozymes were used as markers. Recently, more precise marker techniques such as RFLPs, RAPDs, AFLPs VNTRs, and SSRs were widely used [16]. Each marker technique has its advantages and disadvantages. The choice of a molecular marker technique depends on its reproducibility and simplicity. DNA markers are used to evaluate the genetic variation in gene banks, as well as to identify the phylogenetic and molecular structure of crops and their associated wild species.
The RAPD technique
PCR-based marker like Random Amplified Polymorphic (DNA) (RAPD) was developed independently [17,18]. This technique was widely used to fingerprint a wide range of species from microorganisms to humans. In plants, the technique is used to identify Wheat, Maize, Date, Palms, Corn, and soybeans. cultivars The appeal of this technique was largely due to its ease in applications. However, the random nature of the primers makes the results non-reproducible. The advantages of this technique are:- The markers used seem to be virtually unlimited and are present in all organisms.- RAPD markers have a great potential to detect DNA Polymorphism between genotypes of the same species. DNA (RAPD) is quick and well adapted for nonradioactive DNA fingerprinting of genotypes [19-21]. However, problems with reproducibility in the amplification of RAPD markers and with data scoring have been reported [22]. Although major bands from RAPD reaction are highly reproducible, minor bands can be difficult to repeat due to the random priming nature of this PCR reaction and potential confounding effects associated with co-migration with other markers used 5 RAPD primers to fingerprint and detect the relationships among 17 sorghum cultivars, 11 Egyptian cultivars and 6 Yemenian cultivars [23]. They reported that a total number of 34 polymorphic bands with a level of (94%) polymorphism succeeded to discriminate the sorghum cultivars assayed (51) RAPD primers to determine the genetic distance among 8 Egyptian sweet sorghum cultivars. They found that 49 primers generated a total number of 314 bands which were not necessarily present in each cultivar. 206 bands were found to be polymorphic and (108) bands were common in all of the eight cultivars. The dendrogram tree generated across the 49 primers divided the 8 cultivars into four clusters. The data showed different levels of variation among these cultivars [24,25].
The SSR technique
Simple Sequence Repeats (SSRs) or microsatellites generate a large number of restriction fragments (bands) facilitating the detection of polymorphism which is based on the number of repeat units in a defined region of the genome under investigation [26]. SSRs are ubiquitous in eukaryotes. The number and composition of microsatellite repeats differ in plants and animals. The frequency of repeats longer than 20 bp. has been estimated to occur every 33 kb in plants, unlike mammals where it is found to occur every 6 kb [27]. In humans, (AC) or (TC) are very common repeat units, but in plants, AT is more common followed by AG or TC [28]. In general, plants have 10 times fewer SSRs than humans [29] .Noted that the common SSRs in sorghum are: AG, AC, AAC, and AAG. SSR markers are attractive for DNA fingerprinting studies for several reasons. They are co-dominant and highly informative, occur in high frequency, and appear to be distributed throughout the genomes of most if not all of the higher plants and animals. SSRs generally, display high levels of polymorphism even among closely related accessions and are amenable to automated genotyping strategies. They also can be amplified by inexpensive PCR assays. Radioisotopes are not required in the detection of SSR markers because sequence polymorphism can usually be detected by separation. Most Sorghum bicolor L. SSR loci showed a high level of polymorphism [30].
Materials and Methods
Preparation of plant materials
Seeds of 7 sweet sorghum (Ankolib) accessions, namely, Black Ankolib, Black White Ankolib, Dark Red Ankolib, Red Yellow Ankolib, White, White Black Ankolib, and Bengaga, were collected by the author from three different sites in Sudan. Names of accessions, sites of collection, glume color, and seed color have been shown in Table 2. Similarly, the glume colors for the accessions have been indicated. The seed color of the accession Bengaga is illustrated (Figures 1-7).
TABLE 2 Names of Sweet Sorghum (Ankolib) accessions, site of collection, glume, and seed color.
| No. | Name of Accession | Site | Glume color | Seed color |
|---|---|---|---|---|
| 1 | Black Ankolib | Sinnar | Black | Brown |
| 2 | B.W. Ankolib | Sinnar | Black & White | Brown & Creamy |
| 3 | D.R. Ankolib | Sinnar | Dark Red | Creamy |
| 4 | R.Y. Ankolib | Gadarif | Red & Yellow | Brown & Creamy |
| 5 | White Ankolib | Neyala | White | Creamy |
| 6 | W.B. Ankolib | Gadarif | White & Black | Brown & Creamy |
| 7 | Bengaga | Sinnar | Creamy | Orange |
Preparation of solutions
• Extraction buffer: consist of 2% CTAB (w/v), 100 mm. Tris pH (8.0), (20 mM EDTA (pH 8.0), (1.4) m. NaCl and (0.03%)β-mercaptoethanol (50µl/sample)
• Purification solutions: Two solutions were used in this respect 50 µl Sodium acetate and 1 ml. ethanol absolute and Chloroform: Isoamyl alcohol (CI) 24:13-TE buffer: composed of 10 mm. Tris pH 8.0 and 1 mm. EDTA pH 8.0.
Seed germination
Seeds of each accession were germinated in pots in the lab and left to grow for a week.
DNA Extraction
It is based on the hexa-decyltri-methylamonium bromide (CTAB) method to isolate DNA from tissues containing high levels of polysaccharides and secondary metabolites affecting DNA purification. This modified CTAB 2% protocol includes the use of 1.4 M NaCl, 0.03% β-mercaptoethanol, 50 µl sodium acetate, and 1 ml absolute ethanol for DNA extraction and purification as well as reducing the centrifugation times during the separation and precipitation of the DNA. The tubes were centrifuged at 10000 rpm for 15 minutes to precipitate DNA pellets. The solution in each tube was carefully poured off and the DNA pellets were washed again with 70% ethanol and centrifuged at 10000 rpm for 2 minutes. The solution was poured off and the pellets were air-dried. 200 µl TE buffer or D.D.W. was added and the tubes were left at room temperature for 30 minutes to dissolve the DNA pellets and were then kept at 20ÃÂ??C.DNA purity was determined by a UV-VIS spectrophotometer at a wavelength of 260 nm. The purity was then calculated as the ratio of absorbance at 260 nm to 280 nm.
Concentration of DNA
The DNA was diluted by 1:5 in D.D.W. the samples were electrophoresed in (1%) agarose gel against (0.1 ng) of a DNA size marker (Lambda DNA digested with Hind III and PHIX 174 DNA digested with Hae III). This marker covers a range of DNA fragments size between (23130 bp. and 310 bp.), and a range of concentration between 95 ng and 11 ng (Figure 5). The DNA concentration in a given sample was then estimated by comparing the degree of fluorescence of the unknown DNA band with the different bands in the DNA size marker.
DNA purification
Pure DNA was obtained according to the following procedure: 2 µl RNase was added to each Eppendorf tube and the tubes were transferred to Eppendorf thermomixer 5436 (37ÃÂ??C) for 60 minutes. 50 µl sodium acetate and 1 ml absolute ethanol were added to each tube and the tubes were gently inverted and left for 10 minutes and left in the refrigerator.
PCR reaction
The PCR reaction comprises three steps: denaturation, annealing and extension. The procedure adopted with some modifications.
RAPD-PCR technique
A program was set up and entered in the PCR apparatus to carry out the above-mentioned steps using RAPD primers for 40 cycles as follows: denaturation at 94ÃÂ??C for 5 minutes, annealing at 36ÃÂ??C to amplify DNA for (45 seconds/cycle) and finally extension at 72ÃÂ??C for 1.5 minutes. Another cycle of the extension was conducted at 72ÃÂ??C for 7 minutes.
Preparation of samples for RAPD-PCR reaction
The samples contained a master mix of DNA. Master Mix consisted of 2.5 µl 10X Taq Buffer, 2.5 µl d NTPs 2.5 mm, 2 µl MgCl2 , 25 mm, 2 µl Primer 10 pmol, 0.3 µl Taq DNA Polymerase 5 U/µl and 13.7 µl D.D.W. The 24 RAPD 10-mer primers used in this study were listed in Table 3. 23 µl of the master mix were put in 0.2 ml eppendorf tube and 2 µl template DNA 10 ng was added. The process was repeated 24 times for each DNA sample according to the primer’s number. The tubes were transferred to the PCR apparatus and the program was run. The RAPD-PCR products were stored in a refrigerator at -20ÃÂ??C.
TABLE 3 Names of Sweet Sorghum (Ankolib) accessions, site of collection, glume, and seed color
| No. | Code | Sequence |
|---|---|---|
| 1 | OP-A01 | 5ÃÂ?ÂÂ CAGGCCCTTC 3ÃÂ?ÂÂ |
| 2 | OP-A02 | 5ÃÂ?ÂÂ TGCCGAGCTG 3ÃÂ?ÂÂ |
| 3 | OP-A07 | 5ÃÂ?ÂÂ GTGACGTAGG 3ÃÂ?ÂÂ |
| 4 | OP-A08 | 5ÃÂ?ÂÂ GTGACGTAGG 3ÃÂ?ÂÂ |
| 5 | OP-A09 | 5ÃÂ?ÂÂ GGGTAACGCC 3ÃÂ?ÂÂ |
| 6 | OP-A10 | 5ÃÂ?ÂÂ GTGATCGCAC 3ÃÂ?ÂÂ |
| 7 | OP-A12 | 5ÃÂ?ÂÂ TCGGCGATAG 3ÃÂ?ÂÂ |
| 8 | OP-A15 | 5ÃÂ?ÂÂ TTCCGAACCC 3ÃÂ?ÂÂ |
| 9 | OP-B02 | 5ÃÂ?ÂÂ TGATCCCTGG 3ÃÂ?ÂÂ |
| 10 | OP-B05 | 5ÃÂ?ÂÂ TGCGCCCTTC 3ÃÂ?ÂÂ |
| 11 | OP-B07 | 5ÃÂ?ÂÂ GGTGACGCAG 3ÃÂ?ÂÂ |
| 12 | OP-B12 | 5ÃÂ?ÂÂ CCTTGACGCA 3ÃÂ?ÂÂ |
| 13 | OP-B14 | 5ÃÂ?ÂÂ TCCGCTCTGG 3ÃÂ?ÂÂ |
| 14 | OP-B17 | 5ÃÂ?ÂÂ AGGGAACGAG 3ÃÂ?ÂÂ |
| 15 | OP-B18 | 5ÃÂ?ÂÂ CCCACGCAGT 3ÃÂ?ÂÂ |
| 16 | OP-B19 | 5ÃÂ?ÂÂ ACCCCCGAAG 3ÃÂ?ÂÂ |
| 17 | OP-C02 | 5ÃÂ?ÂÂ GTGAGGCGTC 3ÃÂ?ÂÂ |
| 18 | OP-C20 | 5ÃÂ?ÂÂ ACTTCGCCAC 3ÃÂ?ÂÂ |
| 19 | OP-G02 | 5ÃÂ?ÂÂ GGCACTGAGG 3ÃÂ?ÂÂ |
| 20 | OP-G03 | 5ÃÂ?ÂÂ GAGCCCTCCA 3ÃÂ?ÂÂ |
| 21 | OP-G04 | 5ÃÂ?ÂÂ AGCGTGTCTG 3ÃÂ?ÂÂ |
| 22 | OP-G05 | 5ÃÂ?ÂÂ CTGAGACGGA 3ÃÂ?ÂÂ |
| 23 | OP-G07 | 5ÃÂ?ÂÂ GAACCTGCGG 3ÃÂ?ÂÂ |
| 24 | OP-G09 | 5ÃÂ?ÂÂ CTGACGTCAC 3ÃÂ?ÂÂ |
SSR -PCR Technique
Two programs were adopted here:
• “Touchdown-PCR” adopted from Senior and with minor modifications. SSR primers for this program have been shown in Table 4. 5 SSR primers were screened for amplification of a specific product from sweet sorghum genomic DNA using a (65ÃÂ??C) “touchdown-PCR” technique: denaturation at 94ÃÂ??C for 5 minutes, annealing for 40 cycles, the annealing temperature for the initial 10 cycles was decreased by one degree every cycle from (65ÃÂ??C), followed by 30 cycles at 55ÃÂ??C and extension at (72ÃÂ??C for 1.5 minutes) followed by one more cycle of extension for 7 minutes (Table 4).
TABLE 4 Names of Sweet Sorghum (Ankolib) accessions, site of collection, glume, and seed color.
| Code | Primer ID | Primer Sequence |
|---|---|---|
| A | ZM-ADH2N | F:TGCGAAGAAAGCAGTAGCAAA |
| R:TGGAGGTAGAAGACGCACG | ||
| B | ZM-GPCI | F:CCTCTACTCGCCAGTCGC |
| R:TTTGGTCAGATTTGAGCACG | ||
| C | MZE-ZEINP | F:GGATGATGGACGTGCAGTC |
| R:CTGGTACTGGTAGAGTCCACCC | ||
| S2 | P-bnlg198 | F:GTTTGGTCTTGCTGAAAAATAAAA |
| R: GCTGGAGGCCTACATTATTATCTC | ||
| S11 | P-bnlg615 | F: CTTCCCTCTCCCCATCTCCTTTCCAA |
| R:GCAACCTGTCCATTCTCACCAGAGGATT |
• This program was performed for each pair of the SSR primers present according to the annealing temperature of each primer. Sample according to the SSR primers used in the study. The tubes were placed in the PCR machine and the program was run. The PCR products were kept at (-20ÃÂ??C).
Electrophoresis
Preparation of Running Buffer
This buffer was composed of 50 ml. TBE 10X( pH 8.0) [108g Tris, (55 grams) Boric Acid and (7.44 grams) EDTA in 1 L D.W.] dissolved in (950 ml.) D.W.
Preparation of agarose gel (1.5%)
1.5 grams of agarose gel were added to (100 ml.) TBE running buffer in 250 ml conical flask. The flask was shaken to dissolve the agarose gel, heated in a microwave for about 2 minutes, and left to cool down at room temperature. 3 µl ethidium bromide was added to the cold melted gel, gently shaken, and then poured into a gel tray with a comb to form the wells where The PCR products were loaded and run.
Preparation of acrylamide gel (8%)
The gel was prepared by mixing the following materials:
6.7 ml. acrylamide,15.7 ml D.W., 175 µl TBE 10X, 12.5 TEMED, and 10%APS.
Preparation of samples for running
(5 µl) simple dye (loading dye) was added to each PCR product. Two markers were used to detect polymorphism among PCR products. Lambda X174 RF Hae III marker was used for RAPD-PCR products to determine the molecular weight of bands run in 1.5% agarose gel. 100 bp.
molecular weight of bands run in 1.5% agarose gel. 100 bp. The gels were examined under UV light and were then photographed using a Polaroid Camera and Gel Doc.
Data analysis
The banding patterns generated by Basic-PCR markers RAPD and SSRs were compared to determine the genetic relatedness of the 97) sweet sorghum (Ankolib) accessions. Clear and distinct amplification products were scored as (1) for the presence and (0) for the absence of bands. Bands of the same mobility were scored as identical. The Genetic Similarity Coefficient (GS) between any two genotypes was estimated according to Dice Coefficient as follows:
Dice formula: GSij=2a/(2a+b+c)
Where GSij =the measure of genetic similarity between two individuals i and j.
a = the number of bands shared by i and j.
b = the number of bands present in i and absent in j.
c = the number of bands present in j and absent in i.
Unweighted Pair Group Method using Arithmetic Average (UPGMA) was used for cluster analysis. In this method, each accession represents an independent cluster and the distance between clusters was estimated by the Dice Coefficient. However, once several accessions have been linked together, the distance between two clusters is calculated as the average distance between all pairs of accessions in the two different clusters (Figures 8-10).
Polymorphism detected by SSR markers
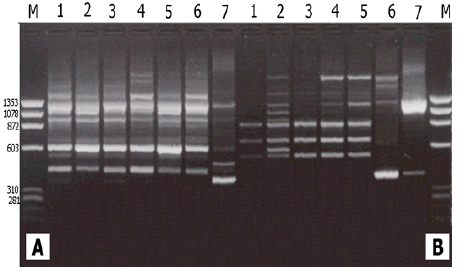
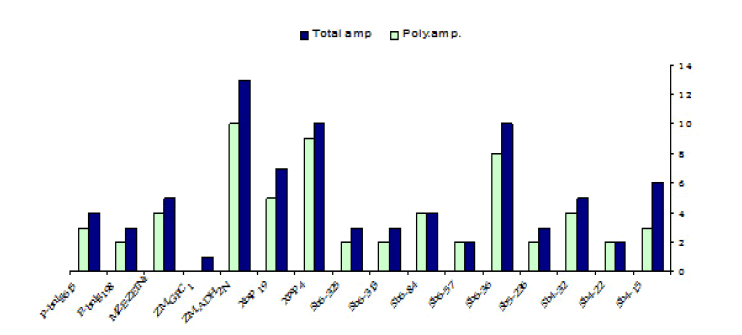
The sixteen SSR primer pairs used in this study successfully generated a total number of 81 bands. One pair (ZM-GPCI) generated only one monomorphic band. The total number of monomorphic fragments detected by the other (15) SSR primer pairs was (19) ranging from (1) to( 3) bands per primer. The total number of polymorphic bands was (62) ranging from (2) to (10) fragments per primer. This represented a level of polymorphism of (76%) and an average of (3.9) fragments per primer. The size of the amplified fragment also varied with the type of primer and it ranged from (50 bp. to 1,100 bp.) (Table 5).
TABLE 5 Primers sequence, the total number of amplicons (amp.), monomorphic amplicons, polymorphic amplicons, and percentage of polymorphism as revealed by SSR analysis
| Primers | F: Forward and R: Reverse sequences | Total amp. | Mono. amp. | Poly. amp. | Poly.% |
|---|---|---|---|---|---|
| Sb4-15 | F:GCTGCTAAGCCGTGCTGA | 6 | 3 | 3 | 50% |
| R:TTATTTGGGTGAAGTAGAGGTGAACA | |||||
| Sb4-22 | F: TGAGCCGAAAACCGTGAG | 2 | - | 2 | 100% |
| R: CCCAAAACCAAGAGGGAAGG | |||||
| Sb4-32 | F: CTCGGCGGTTAGCACAGTCAC | 5 | 1 | 4 | 80% |
| R: GCCCATAGACAGACAGCAAAGCC | |||||
| Sb5-236 | F: GCCAAGAGAAACACAAACAA | 3 | 1 | 2 | 60% |
| R: AGCAATGTATTTAGGCAACACA | |||||
| Sb6-36 | F: GCATAATGACGGCGTGCTC | 10 | 2 | 8 | 80% |
| R: CTTCCAAGTGAAAGAAACCATCA | |||||
| Sb6-57 | F: ACAGGGCTTTAGGGAAATCG | 2 | - | 2 | 100% |
| R: CCATCACCGTCGGCATCT | |||||
| Sb6-84 | F: CGCTCTCGGGATGAATGA | 4 | - | 4 | 100% |
| R: TAACGGACCACTAACAAATGATT | |||||
| Sb6-313 | F: TTCTTCAGTTCGCACAGCATAA | 3 | 1 | 2 | 60% |
| R: ACCTGCAGTGCACTTGACTATTG | |||||
| Sb6-325 | F: AGCGCAGGAGCGCGAA | 3 | 1 | 2 | 60% |
| R:TCATCCGCTACTACCGTACCGTCAGAAA | |||||
| Xtxp4 | F: AATACTAGGTGTCAGGGCTGTG | 10 | 1 | 9 | 90% |
| R: ATGTAACCGCAACAACCAAG | |||||
| Xtxp19 | F: CTTTAATCGGTTCCAGAC | 7 | 2 | 5 | 71% |
| R: CTTCCACCTCCGTACTC | |||||
| ZM-ADH2N | F:TGCGAAGAAAGCAGTAGCAAA | 13 | 3 | 10 | 76% |
| R: TGGAGGTAGAAGACGCACG | |||||
| ZM-GPCI | F: CCTCTACTCGCCAGTCGC | 1 | 1 | - | 0% |
| R: TTTGGTCAGATTTGAGCACG | |||||
| MZE-ZEINP | F:GGATGATGGACGTGCAGTC | 5 | 1 | 4 | 80% |
| R:CTGGTACTGGTAGAGTCCACCC | |||||
| P-bnlg198 | F: GTTTGGTCTTGCTGAAAAATAAAA | 3 | 1 | 2 | 60% |
| R: GCTGGAGGCCTACATTATTATCTC | |||||
| P-bnlg615 | F: CTTCCCTCTCCCCATCTCCTTTCCAA | 4 | 1 | 3 | 75% |
| R:GCAACCTGTCCATTCTCACCAGAGGATT | |||||
| Total | 81 | 19 | 62 | ||
| Average | 5.1 | 1.2 | 3.9 | 76% |
Identification of accessions using unique markers
Unique markers are defined as bands that specifically identify an accession from another by their presence or absence. The bands that are present in one accession but not found in other accessions are termed Positive Unique Markers (PUM), in contrast to the negative unique markers (NUM), which are absent in a specific genotype. These bands are useful for cultivar identification and fingerprinting
Accessions identification by unique RAPD markers
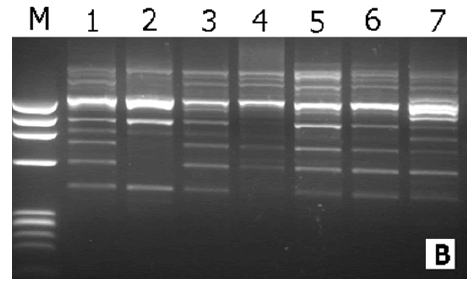
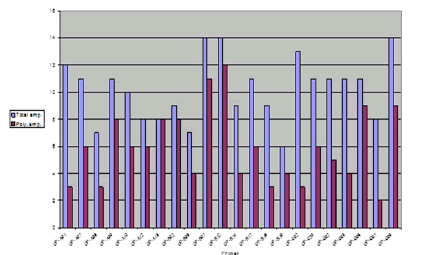
In this study, the RAPD technique was successful in characterizing all of the (7) tested (Ankolib) accessions by unique positive and/or negative markers. These markers ranged in size from (275 bp to 3,000 bp.). A total number of (67) unique markers were identified by the (22) RAPD primers used in this study. Out of the 22 primers, 3 primers (OP-A-15, OP-B-19, and OP-C-20) exhibited only one positive unique marker while 4 primers (OP-A-07, OP-B05, OP-C-02 and OP-G-02) showed only one negative unique marker. The accession Bengaga showed the highest number (27) of the unique markers of which 14 were positive and 13 were negative. This number (27) was obtained from (14) different RAPD primers.
Genetic relationships among the (7) Sweet Sorghum Ankolib accessions as revealed by RAPD markers
To examine the genetic relationships among the study samples based on RAPD results, the scored data were analyzed using the Dice Coefficient to compute the genetic similarity matrices. These similarity matrices were then used to generate a dendrogram using the UPGMA method. The estimated genetic similarity matrices among the accessions ranged from (73.7 to 89.5) thus revealing high levels of genetic similarity among the studied accessions. The highest genetic similarity percentage (90%) is between the accessions: Black Ankolib and Black White Ankolib followed by 89% between the accessions Black Ankolib and Dark Red Ankolib (Figure 11).
Results
Genetic relationships among 7 Sweet Sorghum Ankolib accessions as revealed by SSR markers
The scored data obtained from 16 SSR primer pairs were used to determine the genetic similarities among 7 sweet sorghum Ankolib accessions using the Dice Coefficient. The highest genetic similarity matrix (85.2) was observed for the accessions White Black Ankolib and Black White Ankolib. However, the lowest genetic similarity matrix (66.7) was detected for the following pairs of accessions: Bengaga and Black Ankolib, Black Ankolib and White Ankolib, and finally Red Yellow Ankolib and Black White Ankolib.
Cluster analysis of the data revealed by DNA markers
Matrices of genetic similarities between pairs of individuals may be used as a starting point for statistical analysis of clusters. In a cluster, relatively homogenous groups of individuals cluster together hierarchically and this clustering is visually displayed in a dendrogram.
Cluster analysis of the data revealed by RAPD markers
The UPGMA cluster analysis was carried out to provide a graphical presentation (dendrogram) for the genetic distances among (7) Sweet Sorghum Ankolib accessions. The dendrogram was composed of two clusters. The first cluster included only the accession Bengaga while the second cluster comprised the other (6) remaining accessions. The second cluster included two sub-clusters: the first one comprised the accession Red Yellow Ankolib alone whereas the second one was subdivided into two groups. One group contained the accession White Black Ankolib, whereas the other one was divided into two subgroups. The first subgroup contained the Dark Red Ankolib and the White Ankolib, while the other subgroup contained the Black White Ankolib and the Black Ankolib.
Cluster analysis of the data revealed by combined RAPD- SSR markers
UPGMA cluster analysis was carried out to provide a graphical presentation (dendrogram) for the genetic distances among 7 sweet sorghum Ankolib accessions using the combined data from both markers RAPD and SSR. The dendrogram showed two clusters: the first cluster included the accession Bengaga alone. The second cluster included two sub-clusters: the first one comprised the accession Red Yellow Ankolib alone and the second one had two groups. One group comprised the accession White Ankolib, whereas the other one was divided into two subgroups.SGLT2Is and GN
Discussion
Traditionally, genetic resources in sorghum are classified by taxonomists based on morphological traits governed by a relatively small number of genes. On the other hand, important traits, which are related to habitat adaptation and exhibit enormous variability among sorghum germ plasma, are complex and quantitatively inherited [14].
Molecular genetic identification has become a routine tool in the field of Plant Taxonomy. DNA-based fingerprinting technologies have been proven a useful tool in genetic similarity and genetic relationship studies. The recently developed PCR-based marker techniques, which include RAPD and SSRs are playing an increasingly important role in this type of investigation. In this study RAPD and SSRs techniques were applied to assess three parameters: the first parameter is the assessment of the level of polymorphisms among seven sweet sorghum Ankolib Sorghum bicolor (L.) Moench. accessions collected from three sites in Sudan namely (Sinnar, Gadarif, and Neyala). The second parameter is the identification of unique markers and generating a DNA fingerprint for each accession. The third parameter is the estimation of the genetic relationships among the genotypes
From the results, it was found that each of the two DNA-based markers, RAPD and SSRs, succeeded in detecting a level of polymorphism of (58%) and (76%) respectively. loci are sufficiently polymorphic and indicated that the level of polymorphism at some loci was high enough to allow the vast majority of S. bicolor accessions to be distinguished from one another. This is particularly useful in determining the genotypes at a small number of loci, perhaps as few as half a dozen. This finding is also confirmed by several studies [29,30]. The increased resolution of acrylamide gel over agarose gel could enhance the separation and detection of a larger number of alleles per locus than agarose gel. This may be particularly important for SSR loci containing dinucleotide repeats since PCR products differing by two base pairs cannot be resolved by agarose gels alone. That is why in the present study, we re-electrophoresed all the SSR-PCR products using 8% acrylamide gels to get a more precise estimation of bands.
All of the (7) Ankolib accessions were fingerprinted and uniquely identified by the (22) RAPD primers, which produced a total number of (234) bands of which (138) bands were found to be polymorphic. The (22) RAPD primers gave several unique positive and /or negative fingerprints used (40) RAPD primers of which also (22) primers detected unique positive and/or negative fingerprints [15]. These fingerprints were used in the identification of (15) maize inbred lines. Similarly, a total number of (314) bands was reported by [25] of which (206) were found to be polymorphic. They managed to discriminate 8 Egyptian sweet sorghum cultivars using the (206) polymorphic bands. Of the (16) SSR primers used in this study, only (11) could generate unique positive and/or negative markers. Nevertheless, the banding profiles obtained by the different types of primers yielded unique fingerprints which are characteristic for each accession. The highest number of unique markers was reported for the accession Bengaga using RAPD and SSR markers. It was observed in this study that the accession Bengaga was the only accession that provides grains that could produce flour besides its succulent sweet stem.
The genetic relationships among the (7) sweet sorghum Ankolib accessions were estimated in terms of the genetic similarity matrix using the Dice Coefficient. The ranges of the genetic similarity matrices were (73.7 to 89.5, 66.7 to 85.2, and 72.8 to 86.7) for RAPD, SSRs and RAPDSSRs combined data respectively. These ranges reveal a high level of genetic similarity among the studied accessions thus indicating a high degree of homology.
The highest genetic similarity matrix (89.5) was recorded between accessions Black Ankolib and Black White Ankolib, indicating that these two accessions were closely related to each other. However, the accession Bengaga and Red Yellow Ankolib showed a relatively lower value of the genetic similarity matrix (73.7). This finding agrees with that of those who assessed the genetic diversity among 8 Egyptian sweet sorghum using RAPD markers [25]. The SSR analysis revealed that the highest genetic similarity matrix (85.2) was found between Black White Ankolib and White Black Ankolib. Similarly, the lowest genetic similarity matrix (66.7) was recorded between each of the following pairs of accessions: Black Ankolib & Bengaga, Black Ankolib & White Ankolib, and Red Yellow Ankolib & Black White Ankolib. This indicates the close genetic similarity among the latter accessions.
Cluster analysis of the data obtained by RAPD and SSRs and RAPDSSRs combined data, revealed that the accession Bengaga was a unique cluster as compared to the other (6) (Ankolib) accessions whose seeds were usually smaller than Bengaga and too bitter to be used as flour.
The results obtained from the RAPD technique indicated that the two accessions Black Ankolib and Black White Ankolib lay in the same subgroup.
The RAPD-SSRs combined data revealed that the two accessions belonged to the same division. This reflects the close genetic relationship between the two accessions, a finding which matches the geographical origin (Sinnar Locality) of the accessions. The dendrogram of SSR analysis revealed that the accessions White Ankolib (Neyala) and Red Yellow Ankolib (Gadarif) belonged to the same sub-cluster. Similarly, the two accessions White Black Ankolib (Gadarif) and Black White Ankolib (Sinnar) were found to belong to the same subgroup.
Conclusion
From the above discussion, it can be concluded that DNA-based markers are useful tools for the identification, fingerprinting, and assessment of genetic relationships of sweet sorghum. This study can be summarized as follows:
• The 7 sweet sorghum Ankolib [Sorghum bicolor (L) Moench.] accessions were successfully characterized and uniquely fingerprinted by two DNA-based markers (RAPD and SSRs).
• The accession Bengaga was found to be the most distinct one among all of the 7 accessions in this study.
• There was a close genetic relationship among the other 6 accessions.
• Both SSR and RAPD markers have unprecedented utility for the analysis of population genetics and phylogenetic diversity of sweet sorghum since no radioactive isotopes are required. These markers can be efficiently used by researchers in developing countries.
• The use of SSRs potentially could remove most, if not all of the limitations in revealing polymorphism and in obtaining more complete genomic coverage for plants, as has been achieved already for the human genome.
• Breeders can use genetic similarity information to make informed decisions regarding the choice of genotypes to cross for the development of the sweet sorghum population or facilitate the identification of diverse parents.
• Molecular genetic characterization for some loci controlling glume and seed color in sweet sorghum.
• The use of genetic engineering to develop new varieties of sweet sorghum with high sucrose content and grains for the production of both ethanol and grain flour.
This molecular genetic identification study on sweet sorghum would provide basic molecular genetic data for its genetic engineering in Sudan. Sweet sorghum is a very high biomass-yielding crop. It can be grown as a grain crop, feed crop, sugar crop, and energy crop for making ethanol and electricity. Using ethanol as fuel instead of petrol, will not only save petrol but also can decrease environmental pollution.
References
- Morell MK, Peakall R, Appels R, et al. DNA profiling techniques for plant variety identification. Aust J Exp Agric. 1995;35(6):807-19.
- Rajvanshi AK. Sweet sorghum R&D at the Nimbkar Agricultural Research Institute (NARI). PO. Box. 2005;44.
- Schaffert RE. Sweet sorghum substrate for industrial alcohol. Patancheru AP. 1992;502(324):131-7.
- Koppen S, Reinhardt G, Gartner S. Assessment of energy and greenhouse gas inventories of Sweet Sorghum for first and second generation bioethanol. FAO Environmental and natural resources service series. 2009;30.
- Schaffert RE, Gourley LM. Sorghum as an energy source. In: INTERNATIONAL SYMPOSIUM ON SORGHUM, 1981, Patancheru, India. Sorghum in the eighties: proceedings. Patancheru: ICRISAT. 1982: 605-623.
- Tao Y, Manners JM, Ludlow MM, et al. Henzell RG. DNA polymorphisms in grain sorghum (Sorghum bicolor (L.) Moench). Theoretical and Applied Genetics. 1993;86(6):679-88.
- Zidani S, Ferchichi A, Chaieb M. Genomic DNA extraction method from pearl millet (Pennisetum glaucum) leaves. African Journal of Biotechnology. 2005;4(8):862-6.
- Porebski S, Bailey LG, Baum BR. Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant molecular biology reporter. 1997;15(1):8-15.
- Hoisington D, Khairallah M, Reeves T, et al. Plant genetic resources: what can they contribute toward increased crop productivity?. Proceedings of the National Academy of Sciences. 1999;96(11):5937-43.
- Smith OS, Smith JS, Bowen SL, et al. Similarities among a group of elite maize inbreds as measured by pedigree, F1 grain yield, grain yield, heterosis, and RFLPs. Theoretical and Applied Genetics. 1990;80(6):833-40.
- Nienhuis J, Slocum MK, DeVos DA, et al. Genetic similarity among Brassica oleracea L. genotypes as measured by restriction fragment length polymorphisms. Journal of the American Society for Horticultural Science. 1993;118(2):298-303.
- Dos Santos JB, Nienhuis J, Skroch P, et al. Comparison of RAPD and RFLP genetic markers in determining genetic similarity among Brassica oleracea L. genotypes. Theoretical and Applied Genetics. 1994;87(8):909-15.
- Medraoui L, Ater M, Benlhabib O, et al. Evaluation of genetic variability of sorghum (Sorghum bicolor L. Moench) in northwestern Morocco by ISSR and RAPD markers. Comptes Rendus Biologies. 2007;330(11):789-97.
- Khaled KA, El-Sheikh SR, Tawfik YH. Assessment of genetic diversity among eleven sweet sorghum cultivars (Sorghum bicolor L.) under salt stress. Egyptian Journal of Plant Breeding. 2008;12(1):75-85.
- Zorb C, Schmitt S, Neeb A, et al. The biochemical reaction of maize (Zea mays L.) to salt stress is characterized by a mitigation of symptoms and not by a specific adaptation. Plant Science. 2004;167(1):91-100.
- Drew RA. The application of biotechnology to the conservation and improvement of tropical and subtropical fruit species. Seed and Plant Genetic Resources Service, Food and Agriculture Organization of the Unite Nations; 1997.
- Welsh J, McClelland M. Fingerprinting genomes using PCR with arbitrary primers. Nucleic acids research. 1990;18(24):7213-8.
- Williams JG, Kubelik AR, Livak KJ, et al. DNA polymorphisms amplified by arbitrary primers are useful as genetic markers. Nucleic acids research. 1990;18(22):6531-5.
- Colombo C, Second G, Valle TL, et al. Genetic diversity characterization of cassava cultivars (Manihot esculenta Crantz).: I) RAPD markers. Genetics and Molecular Biology. 1998.
- Fahima T, Sun GL, Beharav A, et al. RAPD polymorphism of wild emmer wheat populations, Triticum dicoccoides, in Israel. Theoretical and Applied Genetics. 1999;98(3):434-47.
- Cao W, Scoles G, Hucl P, et al. The use of RAPD analysis to classify Triticum accessions. Theoretical and Applied genetics. 1999;98(3):602-7.
- Jones CJ, Edwards KJ, Van der Wiel C, et al. Reproducibility testing of SSRs by a network of European laboratories. Molecular tools for screening biodiversity: plants and animals. 1998:209-11.
- Tessier C, David J, This P, et al. Optimization of the choice of molecular markers for varietal identification in Vitis vinifera L. Theoretical and Applied Genetics. 1999;98(1):171-7.
- Prakash SP, Biji KR, Gomez SM, et al. Genetic diversity analysis of sorghum (Sorghum bicolor L. Moench) accessions using RAPD markers. Indian Journal of Crop Science. 2006;1(2):109-12.
- Tautz D. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic acids research. 1989;17(16):6463-71.
- Wang Z, Weber JL, Zhong G, et al. Survey of plant short tandem DNA repeats. Theoretical and applied genetics. 1994;88(1):1-6.
- Powell W, Machray GC, Provan J. Polymorphism revealed by simple sequence repeats. Trends in plant science. 1996;1(7):215-22.
- Mohan M, Nair S, Bhagwat A, et al. Genome mapping, molecular markers and marker-assisted selection in crop plants. Molecular breeding. 1997;3(2):87-103.
- Brown SM, Hopkins MS, Mitchell SE, et al. Multiple methods for the identification of polymorphic simple sequence repeats (SSRs) in sorghum [Sorghum bicolor (L.) Moench]. Theoretical and Applied Genetics. 1996;93(1):190-8.
- Beckmann JS, Soller M. Toward a unified approach to genetic mapping of eukaryotes based on sequence tagged microsatellite sites. Bio/technology. 1990;8(10):930-2.