Whole transcriptome analysis identifies upregulated genes and pathways in ductal carcinoma in situ mimicking usual ductal hyperplasia
2 Department of Pathology, New York University Medical Center, New York, NY, USA, Email: serrano@gmail.com
Received: 29-Sep-2018 Accepted Date: Oct 24, 2018; Published: 30-Oct-2018
Citation: Zeng J, Serrano J, Snuderl M, et al. Whole transcriptome analysis identifies upregulated genes and pathways in ductal carcinoma in situ mimicking usual ductal hyperplasia. J Histol Histopathol Res 2018;2(2): 36-8.
This open-access article is distributed under the terms of the Creative Commons Attribution Non-Commercial License (CC BY-NC) (http://creativecommons.org/licenses/by-nc/4.0/), which permits reuse, distribution and reproduction of the article, provided that the original work is properly cited and the reuse is restricted to noncommercial purposes. For commercial reuse, contact reprints@pulsus.com
Abstract
Ductal carcinoma in situ (DCIS) and usual ductal hyperplasia (UDH) represent two extremes on the spectrum of intraductal proliferative lesions of the breast with corresponding differences in invasive carcinoma risk, molecular alteration and therapeutic management. In cases with overlapping morphology, the distinction between DCIS and UDH can be challenging. We performed whole transcriptome analysis on laser-capture microdissected samples from formalin-fixed, paraffin-embedded (FFPE) tissue in two cases of DCIS with UDH-like morphology (DUM) and their corresponding UDH. Of the top 10 genes differentially up/down regulated in DUM compared to UDH, we selected cyclin D1 (upregulated in DUM) and CD117 (c-kit) (down regulated in DUM) and performed immunohistochemical studies using clinically validated antibodies for cyclin D1 and CD117, respectively. Our results showed that both DUM lesions showed strong and diffuse (>50%) nuclear positivity for cyclin D1 antibody. The corresponding foci of UDH showed no reactivity to weak and sparse (<1%) staining for cyclin D1 antibody. Conversely, UDH expressed patchy CD117 (c-kit) positivity, while DUM lesion was completely negative. We also demonstrated that DUM lesions are enriched for genes associated with invasive ducal carcinoma, luminal subtype and brain metastasis. In summary, despite morphological similarities, DUM and UDH are molecularly distinct. Our study showed that whole transcriptome analysis of morphologically distinct lesions from the same FFPE sample isolated by laser capture microdissection provides a powerful tool to identify novel biomarkers in breast cancer.
Keywords
Ductal carcinoma in-situ; Usual ductal hyperplasia; Whole transcriptome analysis
ntraductal proliferative lesions of the breast are classified into three categories: usual ductal hyperplasia (UDH), atypical ductal hyperplasia (ADH) and ductal carcinoma in situ (DCIS). While distinguishing UDH from DCIS is straightforward in the majority of cases, there are cases of DCIS that closely mimic UDH morphology (herein referred to as DCIS with UDH-like morphology or “DUM”). The implications of misdiagnosis cannot be overemphasized. Patients with DCIS often require a combination of surgical excision with negative margins, radiation therapy and endocrine therapy while UDH is not an actionable diagnosis and has traditionally been considered a benign diagnosis with no further therapy. We present high throughput molecular analysis of two cases of DUM with its corresponding UDH to characterize their molecular landscape and identify putative diagnostic markers, which may aid in the diagnosis as a supplement to the existing markers.
Case Report
Two cases of DUM was identified in 2016 and discussed at our peer-review conference due to diagnostic difficulties. Whole transcriptome analysis was performed on each case of DUM with respective foci of UDH, using pure samples obtained from laser capture microdissection (LCM) from five 10-μmthick sections of formalin-fixed, paraffin-embedded (FFPE) tissue. RNASeq libraries were prepared using the Clontech SMARTer Stranded Total RNASeq Kit library prep, following the manufacturer’s protocol. The libraries were loaded on high output HiSeq 2500 flow cells generating 60-80 million reads per sample. Data were analyzed using our in-house bioinformatics pipeline including Database for Annotation, Visualization and Integrated Discovery (DAVID) and Gene Set Enrichment Analysis (GSEA). Immunohistochemical (IHC) studies were performed using estrogen receptor (ER) (SP1, Ventana, Tucson, AZ), progesterone receptor (PR) (1E2, Ventana, Tucson, AZ), cytokeratin 5/6 (CK5/6) (D5/16B4, Ventana, Tucson, AZ), synaptophysin (SP11, Ventana, Tucson, AZ), chromogranin (LK2H10, Ventana, Tucson, AZ), cyclin D1 (SP4R, Ventana, Tucson, AZ) and CD117 (c-kit; polyclonal rabbit, Dako, Carpenteria, CA) antibodies [1-3].
Results
Case 1 is that of a 60-year-old woman, who presented with bloody nipple discharge followed by surgical excision. Microscopic examination showed an aggregate of distended ducts with mostly cribriform, solid and focally papillary architecture populated by ductal cells with round to oval nuclei and blood (Figure 1A).
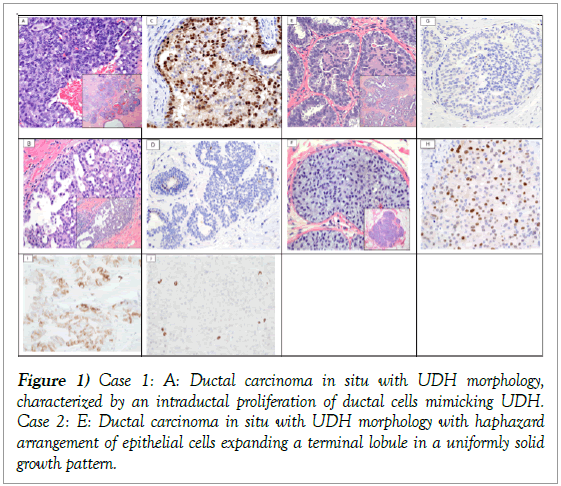
Figure 1: Case 1: A: Ductal carcinoma in situ with UDH morphology, characterized by an intraductal proliferation of ductal cells mimicking UDH. Case 2: E: Ductal carcinoma in situ with UDH morphology with haphazard arrangement of epithelial cells expanding a terminal lobule in a uniformly solid growth pattern.
Case 1: A: Ductal carcinoma in situ with UDH morphology, characterized by an intraductal proliferation of ductal cells mimicking UDH. Inset: At lower magnification, DUM shows distended ducts with irregular lumen formations. (H&E, original magnification 40x; inset at 4x) B: Usual ductal hyperplasia characterized by a heterogeneous population of epithelial cells. Inset: At lower magnification, the intraductal proliferation completely fills the lumen forming bridges, tufts and irregular fenestrations. (H&E, original magnification 40x; inset at 10x) C: Immunohistochemistry for cyclin D1 protein expression in the DUM lesion shows strong and diffuse nuclear reactivity in the majority of luminal cells. (cyclin D1, original magnification 40x) D: Immunohistochemistry for cyclin D1 protein expression in the UDH lesion shows rare cells with nuclear positivity. (cyclin D1, original magnification 40x). The cells typically lacked polarity and housed oval nuclei with somewhat irregular contour and occasional grooves. The nuclei were not equidistant, a feature which imparted an overall “streaming pattern” and “nuclear overlapping” typically seen in UDH. Despite that, the cells were more or less monomorphic with little variation in nuclear size and shape unlike the adjacent UDH (Figure 1B). Case 2 is that of a 44-year-old woman, who had a stereotactic core biopsy for microcalcifications diagnosed as atypical ductal hyperplasia followed by surgical excision. Microscopic examination, similar to case 1, showed scattered distended ducts with solid architecture populated by ductal cells with haphazard and not equidistant cellular arrangement and swirling. The cells showed oval nuclei with dark chromatin without nucleoli. Occasional intranuclear inclusions were seen. There were adjacent foci of UDH with its characteristic morphology (Figures 1E and 1F). Case 2: E: Ductal carcinoma in situ with UDH morphology with haphazard arrangement of epithelial cells expanding a terminal lobule in a uniformly solid growth pattern. (H&E, original magnification 40x; inset at 10x) F: Usual ductal hyperplasia with a heterogeneous population of epithelial cells forming large tufts and incomplete bridges seen at both high and low magnification (inset), an architectural growth pattern that is typical for UDH. (H&E, original magnification 40x; inset at 10x) G: Immunohistochemistry for cyclin D1 protein expression in the UDH lesion shows no to rare and weak nuclear reactivity (cyclin D1, original magnification 40x) H: Immunohistochemistry for cyclin D1 protein expression in the DUM shows diffuse nuclear reactivity in at least 50% of luminal cells with variable intensity. (cyclin D1, original magnification 40x); I: Immunohistochemistry for CD117 (c-kit) shows patchy membranous staining in UDH (CD117, original magnification 40x); J: Immunohistochemistry for CD117 (c-kit) shows no staining of DUM cells. Notice the entrapped mast cells staining for c-kit (CD117; original magnification 40x) (Figure 1). Immunohistochemically, the atypical lesional cells in both cases were diffusely positive for ER and PR and negative for synaptophysin, chromogranin and CK5/6. Based on the cytomorphology, extent and immunoprofile, we made the diagnosis of DCIS in both cases. Given the similarities to UDH, we dubbed the term DUM for the purpose of reporting in the manuscript [4].
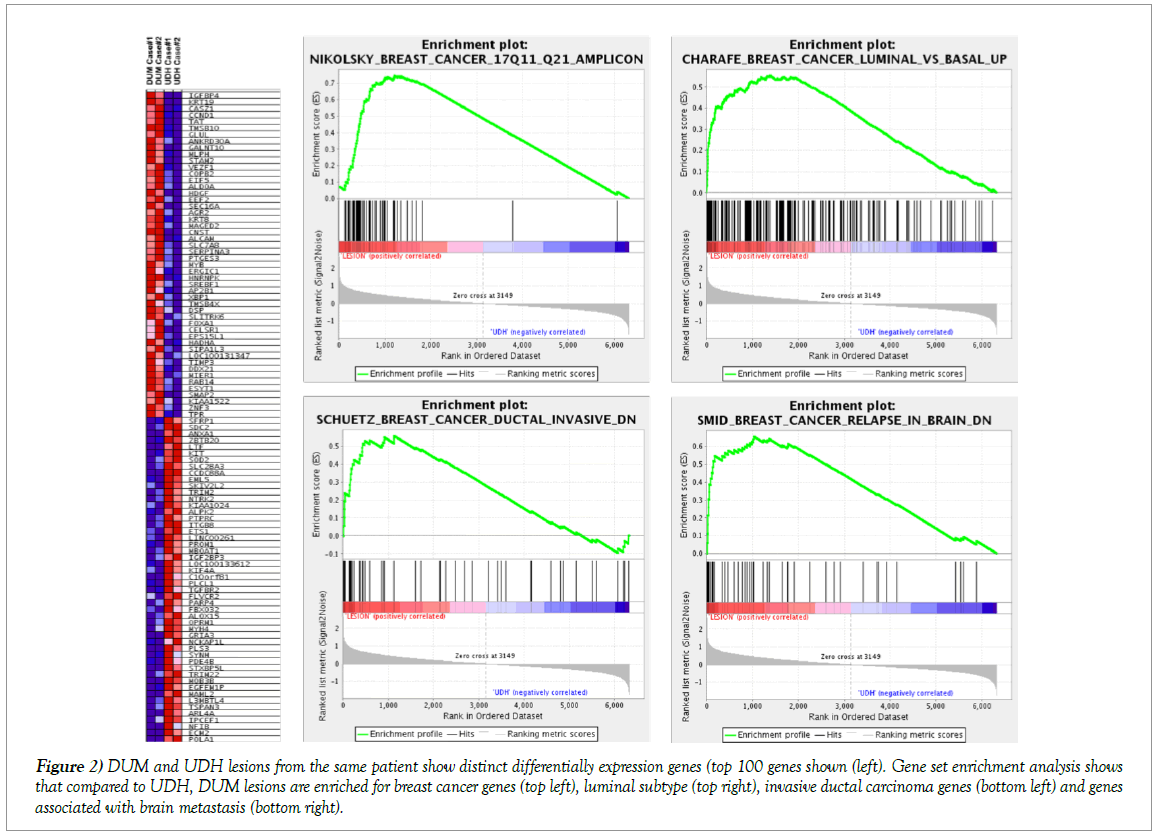
The DUM cases were significantly enriched for genes previously identified as overexpressed in breast cancers and luminal breast cancer and enriched for genes upregulated in breast cancer metastasis to the brain and invasive ductal carcinoma (Figure 2). The top 10 upregulated and downregulated genes in DUM compared to UDH are listed in Figure 2. To confirm differentially up/downregulated genes identified by RNAseq, we selected cyclin D1 (upregulated in DUM) and CD117 (c-kit) (downregulated in DUM) and performed IHC using clinically validated antibodies for cyclin D1 and CD117, respectively. Both DUM lesions showed strong and diffuse (≥50%) nuclear positivity for cyclin D1 antibody (Figures 1C and 1H). The corresponding foci of UDH showed no reactivity to weak and sparse (<1%) staining for cyclin D1 antibody (Figures 1D and 1G). Conversely, UDH expressed patchy CD117 (c-kit) positivity, while DUM lesion was completely negative (Figures 1I and 1J respectively) [5].
Figure 2: DUM and UDH lesions from the same patient show distinct differentially expression genes (top 100 genes shown (left). Gene set enrichment analysis shows that compared to UDH, DUM lesions are enriched for breast cancer genes (top left), luminal subtype (top right), invasive ductal carcinoma genes (bottom left) and genes associated with brain metastasis (bottom right).
Discussion
Whole transcriptome analysis and RNAseq can be successfully performed on small amounts of laser-capture microdissected FFPE tissue to identify differentially expressed genes. The results can be further validated using clinical antibodies to establish novel biomarkers that can improve the diagnosis. In this study, we utilized RNAseq to highlight the distinct molecular signature of DCIS in comparison to its corresponding UDH in two challenging cases. In general, DCIS can be accurately diagnosed and graded by adhering to the established histologic criteria; however, challenging lesions exist. In certain cases of DCIS, especially those with grade 2 nuclei, the cellular arrangement and architecture suggest the diagnosis of UDH. This can lead to diagnostic variability and can adversely influence clinical management. 1-4
The top 10 genes upregulated in DUM are implicated in a variety of cellular functions including cell metabolism (TAT, GLUL, GALNT10), transcription factors (CASZ1, ANKRD30A), organization of cytoskeleton (KRT19, TMSB10) and insulin-like growth factor binding protein family and cell signaling (IGFBP4). Among them is CCND1, which encodes cyclin D1, a cell cycle regulator protein. In the breast, CCND1 is posited to be one of the tumor suppressor genes responsible for the early development of DCIS. 5 We confirmed CCND1 RNA upregulation in DUM at the protein level using anti-cyclin D1 antibody.
Ancillary studies can aid the pathologist in making the correct diagnosis in challenging cases of UDH versus ADH or low grade DCIS. Typically, UDH displays a heterogeneous (mosaic) staining pattern for CK5/6 and ER while ADH and low grade DCIS show lack of staining for CK5/6 and diffuse and strong staining pattern for ER. Our two DUM cases showed staining patterns consistent with the diagnosis of DCIS while the respective UDH foci showed a typical mosaic staining pattern for CK5/6 and only focal positivity for ER. We propose that cyclin D1 and CD117 may be of help as a supplement to the existing markers to help distinguish DCIS from UDH though this proposal needs to be validated by larger series.
Solid-papillary carcinoma is another entity that can morphologically mimic UDH. At least a subset of solid-papillary carcinoma is considered a DCIS variant and a non-obligate precursor to invasive mucinous carcinoma. In these cases, attention to monotony and nuclear palisading of the neoplastic cells, intra- and extracellular mucin, hyalinized fibrovascular cores and neuroendocrine differentiation, as evidenced by immunoreactivity for synaptophysin and chromogranin, in solid-papillary carcinoma can aid in making the correct diagnosis. The two DUM cases were negative for synaptophysin and chromogranin. Also there were no hyalinized fibrovascular cores in the foci of DUM.
In summary, we present molecular analysis of two cases of DUM with their matched UDH. We demonstrate that despite morphological similarities, DUM and UDH are molecularly distinct. We confirmed the upregulation and downregulation of cyclin D1 and CD117 in DUM, respectively, at the RNA and protein level. In addition, we demonstrated that whole transcriptome analysis of morphologically distinct lesions from the same FFPE sample isolated by LCM provides a powerful tool to identify novel biomarkers in breast cancer.
Acknowledgement
Conflicts of Interest and Source of Funding: The corresponding author and remaining authors do not have any conflicts of interests to declare. Ethics approval and consent from our institution was requested, however our IRB do not require approval for reporting individual cases or case series.
REFERENCES
- Page DL, Dupont WD, Rogers LW, et al. Atypical hyperplastic lesions of the female breast. A long-term follow-up study. Cancer. 1985;55(11):2698-708.
- Tavassoli FA, Norris HJ. A comparison of the results of long-term follow-up for atypical intraductal hyperplasia and intraductal hyperplasia of the breast. Cancer. 1990;65(3):518-29.
- Ellis IO. Intraductal proliferative lesions of the breast: morphology, associated risk and molecular biology. Modern pathology: an official journal of the United States and Canadian Academy of Pathology, Inc. 2010;23 Suppl 2:S1-7.
- Mitchell KB, Kuerer H. Ductal Carcinoma In Situ: Treatment Update and Current Trends. Current oncology reports. 2015;17(11):48.
- O'Connell P, Pekkel V, Fuqua SA, et al. Analysis of loss of heterozygosity in 399 premalignant breast lesions at 15 genetic loci. Journal of the National Cancer Institute. 1998;90(9):697-703.