The nucleolar ribosome factory: A coaxial cable at heart
Received: 08-Nov-2021 Accepted Date: Nov 22, 2021; Published: 29-Nov-2021
Citation: Tartakoff A, Lin S.The nucleolar ribosome factory: A coaxial cable at heart. J Histol Histopathol Res.2021;5(4):1-2.
This open-access article is distributed under the terms of the Creative Commons Attribution Non-Commercial License (CC BY-NC) (http://creativecommons.org/licenses/by-nc/4.0/), which permits reuse, distribution and reproduction of the article, provided that the original work is properly cited and the reuse is restricted to noncommercial purposes. For commercial reuse, contact reprints@pulsus.com
Description
As a result of more than a century of cytological investigations, the nucleolus of higher eukaryotic cells has traditionally been thought to consist of three sub compartments/phases that function sequentially in the biogenesis of ribosomal subunits [1,2]. Our studies of the yeast, S.cerevisiae, have led us to realize that the nucleolus in fact consists of a “coaxial cable” in which the rDNA axis is surrounded by two layers (inner, outer) of distinct composition [3]. These layers are composed of the ~ 200 Assembly Factors (AFs) that contribute to the biogenesis of either the Small Subunit (SSU) or Large Subunit (LSU) and account for most of the mass of the corresponding immature particles.
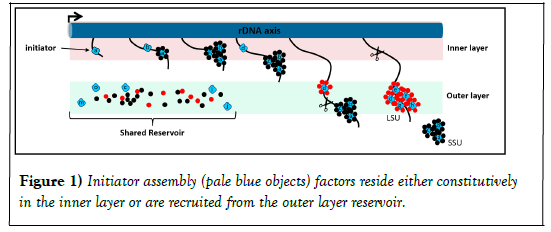
According to this model, when the cell is not producing ribosomal subunits, AFs are in their latent states, and most of the factors are in the outer layer, while a few - that appear to perform initiator functions - reside in the inner layer. We think of the many factors in the outer layer as constituting a shared “reservoir." Once rDNA transcription begins, we propose that the AFs are recruited from the reservoir to the nascent rRNA which is initially close to the axis itself. Due to the linear organization of rDNA, the 5' end of the transcript begins with a spacer (5'-ETS) that is followed by the sequences of SSU rRNA and then LSU rRNAs. Following this order, initiator AFs binds the 5’-most sequences of nascent rRNA in the inner layer and subsequently recruits additional SSU AFs from the reservoir. The result is the production of oversized "knob-like" particles that enclose the maturing subunit. Ribosomal proteins are added throughout this process. As transcription continues, the immature SSU particle extends and actually reaches into the outer layer, where it is released by nuclease cleavage. LSU rRNA sequences then continue to extend, initiator LSU AFs are recruited from the nearby reservoir, and further stages of assembly ensue, roughly as for SSU. When the 3' end of the rRNA has been transcribed, the immature LSU particle is severed and - like the SSU particles - undergoes further surface remodeling as it enters the nucleoplasm. In the "return arm" of this cycle, the released latent assembly factors relocalize to the inner or outer layers, as appropriate (Figure 1).
In both cases they bind specific sites along nascent rRNA transcripts when those emerge. Secondary assembly factors (small circles) are then recruited from the reservoir in the outer layer to bind to these primary units. Their subunit specificity is indicated by color (black for SSU, red for LSU). The scissors indicate the sites of RNA cleavage.
With this model in mind, it is of interest to understand why latent forms of different AFs localize to different layers and especially why and how the SSU assembly intermediates are transferred from the inner to the outer layer where the rRNA is cleaved and subunit precursors are released. We hypothesized that transfer is driven by potential energy relations [3-11]. The basic idea is that the latent SSU AFs found in the outer layer are most stable in that layer and that their association with nascent rRNA recruits them into the environment that prevails in the inner layer, where they are less stable. There therefore is a progressive tendency for them (and the rRNPs in which they reside) to relocate to the outer layer.
But is it reasonable to postulate the existence of a non-subunit-specific shared reservoir? Alternatively, there might be intrinsic differences between the latent forms of the proteins of the two layers. Perhaps latent AFs that contribute to each subunit have some distinct characteristic. For example, differences in charge, 2o/3o structure, relative disorder, or solubility might contribute. Any identification of generic differences might ultimately help rationalize the existence of the two layers and reveal the biochemical basis for their separation.
Conclusion
Earlier studies have shown that the large majority of the AFs in S. cerevisiae can be traced back to the last common ancestor of eukaryotes and that their homologs are also found in higher eukaryotes. We therefore expect that the coaxial model will prove to be universal. Its detection has relied on methods that make it possible to unravel the otherwise overcrowded internal environment of the nucleolus. We anticipate that insight into the functional organization of the nucleolar "ribosome factory" will help elucidate these specific events and also the packaging and processing of other RNAs, both in health and in disease.
It has long been evident that the appearance of the nucleolus is sensitive to many forms of stress, viral infection and malignancy. There may also be a link to age judging from studies of progeria. Moreover, features of ribosomopathies may reflect malfunction of the nucleolus. An in-depth understanding of any of these perturbations will require understanding of normal function.
REFERENCES
- Lafontaine DLJ, Riback JA, Bascetin R, et al. The nucleolus as a multiphase liquid condensate. Nat Rev Mol Cell Biol. 2021;22(3):165-182.
- Pederson T. The nucleolus. Cold Spring Harb Perspect Biol. 2011;3(3):638.
- Tartakoff AM, Chen L, Raghavachari S, et al. The nucleolus as a polarized coaxial cable in which the rDNA axis is surrounded by dynamic subunit-specific phases. Curr Biol. 2021; 31(12):2507-2519.
- Ebersberger I, Simm S, Leisegang MS, et al. The evolution of the ribosome biogenesis pathway from a yeast perspective. Nucleic Acids Res. 2014;42(3):1509-1523.
- Bohnsack KE, Bohnsack MT. Uncovering the assembly pathway of human ribosomes and its emerging links to disease. EMBO J. 2019;38(13):278.
- Badertscher L, Wild T, Montellese C, et al. Genome-wide RNAi Screening Identifies Protein Modules Required for 40S Subunit Synthesis in Human Cells. Cell Rep. 2015;13(12):2879-2891.
- Weeks S, Metge B, Samant R. The nucleolus: a central response hub for the stressors that drive cancer progression. Cell Mol Life Sci. 2019;76(22):4511-4524.
- Montanaro L, Trere D, Derenzzini M. Nucleolus, ribosomes, and cancer. Am J Pathol. 2008;173(2):301-310.
- Buchwalter A, Hetzer M. Nucleolar expansion and elevated protein translation in premature aging. Nat Commun. 2017;8(1):328.
- Boulon S, Westman B, Hutten S,et al. The nucleolus under stress. Mol Cell. 2010;40(2): 216-227.
- Farley-Bownes K, Ogawa L, Baserga S. Ribosomopathies: Old Concepts, New Controversies. Trends Genet. 2019;35(10):754-767.